Emerging trends in human skin microbiota in diabetic and non-diabetic subjects
Received:
Accepted:
Published:
Authors: Windy Cole, Nicholas Butler, Emma Woodmansey, Janina A. Krumbeck, Kaylie Zapanta, Livia Nguyen, Nina Kovolyan, Sierra Clark & Paola Pena
Citation:
Cole W, Butler N, Woodmansey E et al (2025) Emerging trends in human skin microbiota in diabetic and non-diabetic subjects. Global Wound Care Journal 1(2): 8–19
Conflicts of interest:
Funding for this research was supported by Zymo Research Corporation. JAK and KZ are employed by Zymo Research Corporation. WC is on the Global Wound Care Journal editorial board; this did not influence peer review.
Acknowledgements:
The authors extend their gratitude to the following individuals, whose invaluable contributions significantly enhanced this project: Vikram Samant, Porscha Dort, Tobin Huffman and Manpreet Kaur; and Stacey Coe for her support in protocol writing.
Corresponding author:
Dr Windy Cole, Director of Wound Care Research, College of Podiatric Medicine, Kent State University, 6000 Rockside Rd, Independence, OH, USA. Email: wcole4@kent.edu
DOI: https://doi.org/10.63896/gwcj.1.2.8
Introduction: The skin hosts a diverse microbiota, and its disruption can lead to pathogen colonisation. This study aimed to assess differences in skin microbiota between individuals with and without diabetes, and explore variations by sex, skin location and skin tone.
Methods: A cross-sectional study of 100 participants with diverse skin tones (50 diabetic, 50 non-diabetic; 43 males, 57 females) analysed microbiota from the right plantar forefoot and interdigital spaces using next-generation sequencing.
Results: Both groups shared similar bacterial microbiota, with Staphylococcus hominis and S. epidermidis being the most common. However, the fungal diversity differed between groups, and diabetic participants had higher abundances of Gram-negative pathogens, particularly Klebsiella aerogenes pneumoniae and a Pseudomonas sp., in the plantar region. The interdigital spaces of diabetics also showed an increase in a Pseudomonas sp., especially P. aeruginosa, while non-diabetics had more Gram-positive Streptococci. Various positive interactions between commensal and pathogenic microbes differed between groups.
Conclusions: This study highlights microbiota variations associated with diabetes, skin location, and other factors. Whether these changes contribute to or result from diabetes remains unclear. Further research may enhance understanding of skin health and infection risks in diabetes.
Level of clinical evidence: Prospective and retrospective cohort studies (n≥30 patients) = 2
The microbiota of the skin is composed of a complex community of microorganisms, including bacteria and fungi, in which humans coexist as a host (Chen and Tsao, 2013; Skowron et al, 2021; Bay and Ring, 2022). The cutaneous microbiota plays a crucial role in supporting various functions, including immune responses and fortifying the protective barrier of the epidermis (Harris-Tryon and Grice, 2022).
Both host-related and external factors can influence the composition of the microbiota. However, human flora shows a certain amount of resistance to change, with the tendency to recover to baseline after a mild alteration (Cho and Blaser, 2012). The skin is the largest organ of the body and is vulnerable to environmental perturbations. The colonisation of the microorganisms that comprise the skin microbiota varies based on individual host factors and environmental influencers. Typically, the skin is cool, dry and acidic. However, variations in the physical and chemical composition of regions of skin contribute to the adaptation of a unique set of microorganisms that inhabit a particular microenvironment (Grice and Segre, 2011). Skin topography, including thickness, ridges and folds, hair follicles and glandular structures, can significantly alter the microbiota (Tagami, 2008). Additionally, individual, endogenous host factors (e.g. age, gender, race, genetics, pregnancy status, socioeconomic status, environmental exposure and comorbidities) can cause variability in the skin microbial flora (Grice and Segre, 2011; National Academies of Sciences, Engineering, and Medicine, 2018). On comorbidity, hyperglycaemia from uncontrolled diabetes has been linked to bacterial virulence attributed to chronic infection, including tissue adherence and biofilm formation.
Alterations in the cutaneous microbiota can not only disrupt the host–microbe relationship, but may also permit unchecked cutaneous colonisation by pathogens. In response to diabetes-induced microbial alterations, resident microflora members can help to protect the host from infection related to diabetes, shifting potential pathogens to more commensal-like behaviour (Ramsey et al, 2016). However, this area of research is in its infancy. No study to date has investigated the skin microbiota among individuals with diabetes and the comorbidities associated with the disease. As such, this study aimed to explore if there was a detectable difference in the diversity, quantity of organisms found in the skin microbiota of the plantar surfaces and interdigital spaces of the feet of individuals with and without diabetes. Additionally, it was investigated whether trends exist in the microbiota that can explain the development of foot infections and chronic ulcerations related to diabetes.
Patient materials and methods
This research was conducted as a single-centre, prospective, non-randomised cohort study aimed at evaluating the cutaneous microbiota in the plantar forefoot and interdigital spaces. This study received approval from the Institutional Review Board (IRB), emphasising our commitment to ethical standards.
Study participants:
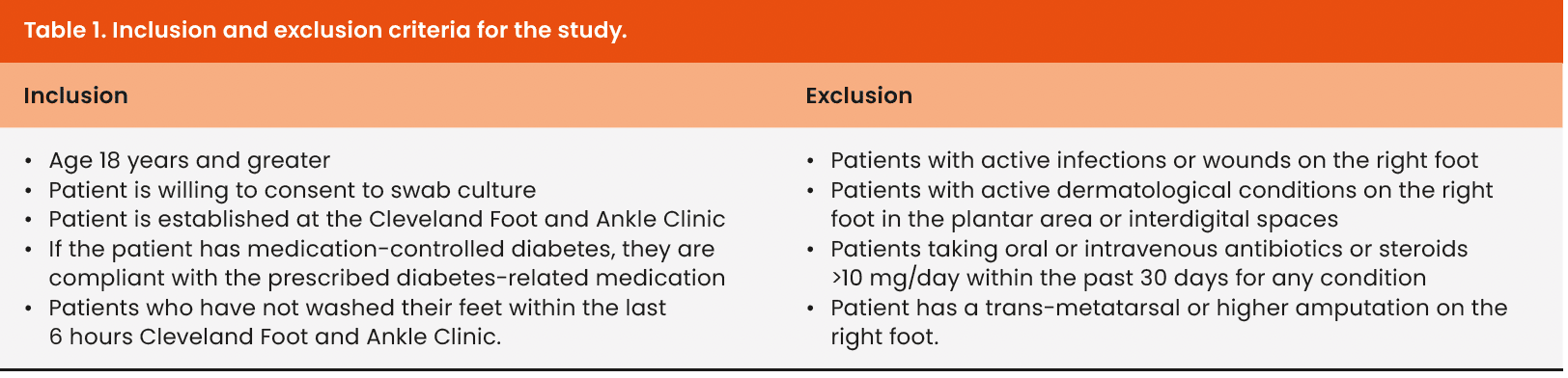
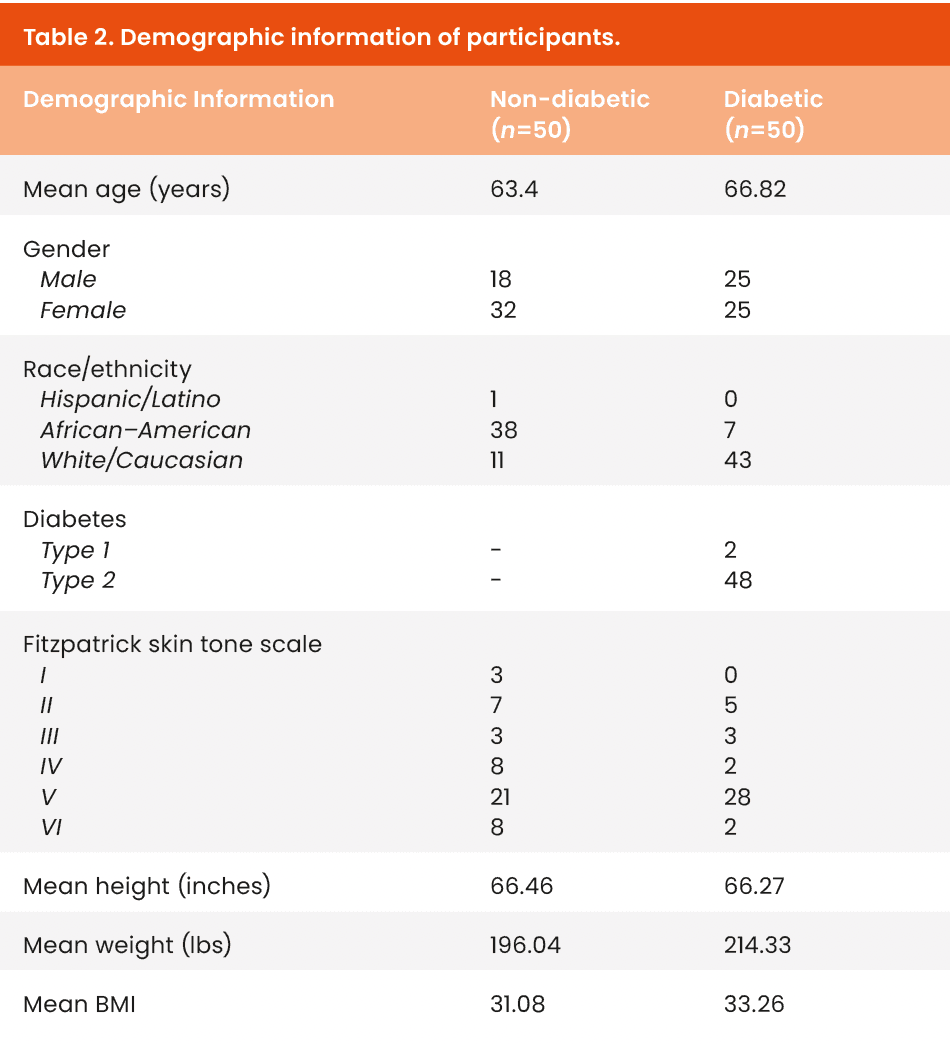
One hundred participants were recruited for this study: individuals with diabetes (n=50) and without diabetes (n=50). Participants in the study were patients at the Cleveland Foot and Ankle Clinic. The detailed inclusion and exclusion criteria can be found in Table 1. Each cohort comprised 50 patients, ensuring a balanced and comprehensive analysis. We obtained written informed consent from each participant in private clinic settings, granting individuals sufficient time to review information, pose questions, and receive thorough answers. We collected demographic data from participants, including age, gender, height, weight, BMI, race, ethnicity, tobacco use and the presence and type of diabetes, along with the Fitzpatrick skin type scale [Table 2] (Tagami, 2008).
Sample collection
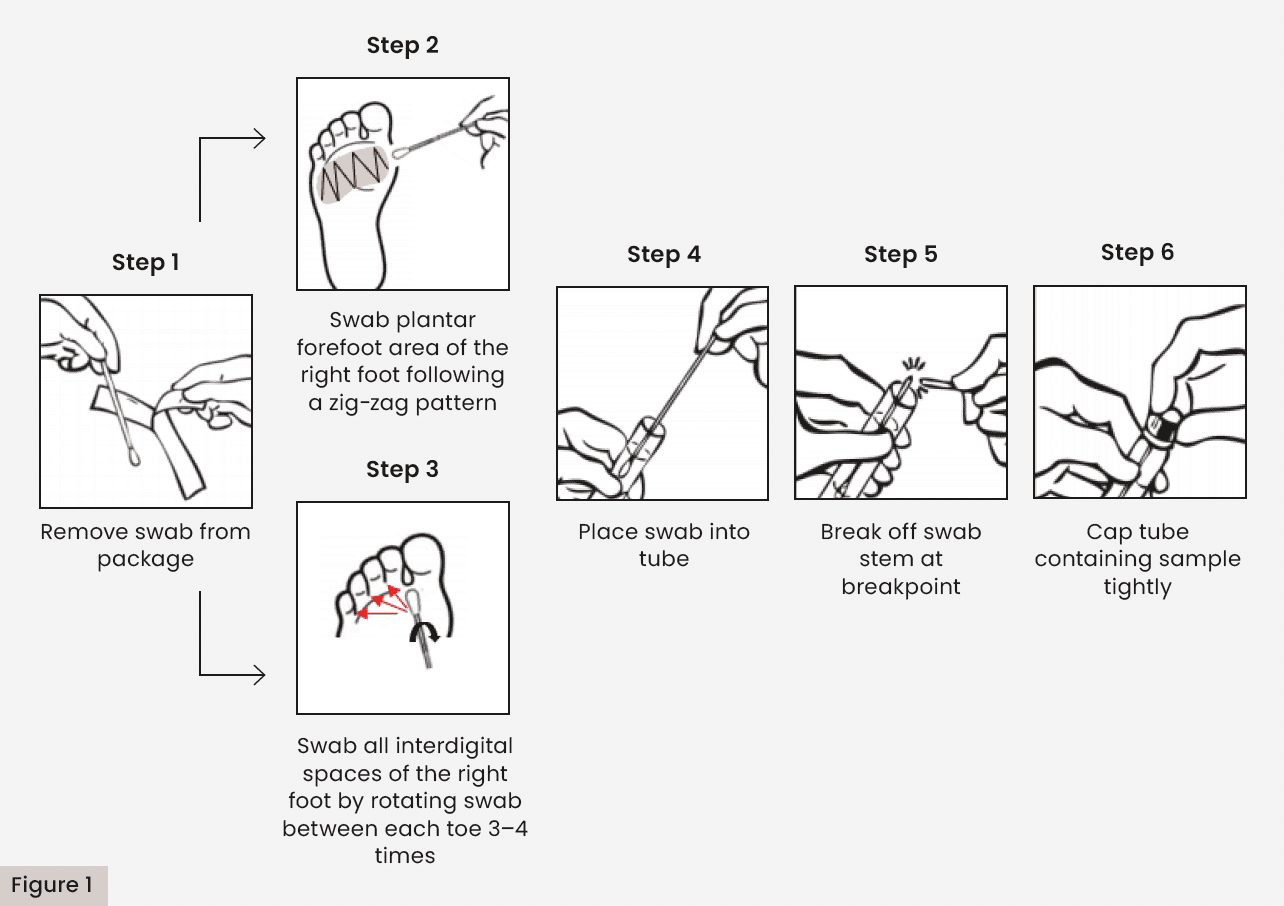
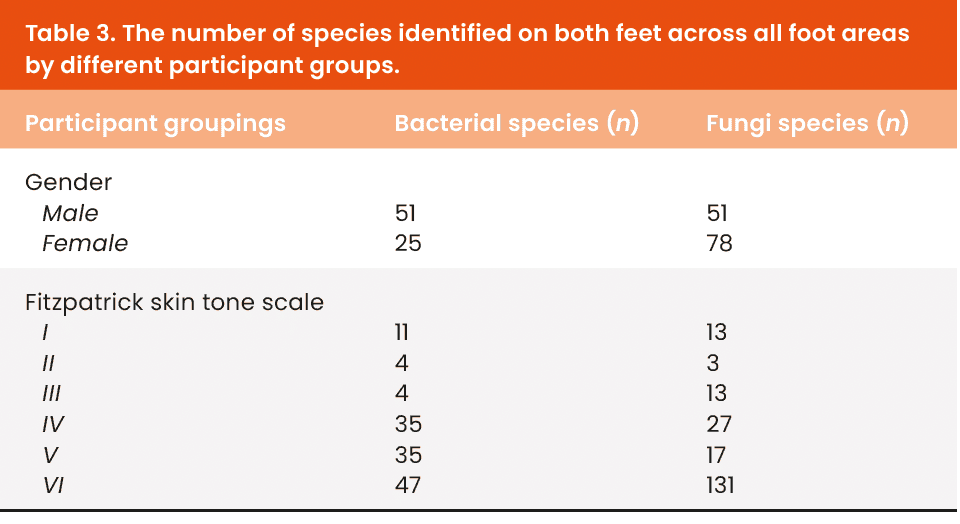
A total of two separate skin swabs were collected from each patient – one from the plantar forefoot and one from the interdigital spaces of each subject’s right foot, as shown in Figure 1. Table 3 shows the number of species identified on both feet across all foot areas by different participant groups. Specimens were collected in kits supplied by Pangea Laboratories (Tustin, CA, USA). Collection devices consisted of a sterile swab and a collection tube pre-filled with 2 ml of DNA/RNA Shield™, a reagent that stabilises and preserves nucleic acids present in the specimen. After swabbing the area of interest, the swab was placed directly into the collection tube. The swab stem was broken off at the breakpoint, and the collection tube was capped tightly to prevent leakage and loss of specimen during shipment. Tubes were labelled with the patient ID and site of collection, plantar forefoot and inter-digital space (IDS).
Sample storage
Properly collected specimens were stable for up to 1 month at room temperature. Once collected, the two collection tubes per patient were placed in a provided biohazard bag. The sample information form specific to that patient was inserted in the bag’s front pocket. Bagged specimens were stored in a secure cabinet in the research office until they were shipped. Pangea Laboratories stored specimens for a limited time (i.e. up to 6 months at −20°C from the time of collection). Samples were kept only for the purpose of this study and were not used for projects outside the scope of this study.
Sample shipping
Specimens were batch shipped to Zymo Research Corporation (Irvine, CA, USA) for processing and next-generation sequencing (NGS) analysis. Bagged specimens were placed into a shipping box and shipped from the Cleveland Foot and Ankle Clinic by International Air Transport Association (IATA) certified study staff using return labels supplied by Pangea Laboratories.
Sample processing
Microbial DNA from the specimen was extracted using the ZymoBIOMICS DNA kit, according to the manufacturer’s instructions, and prepared for NGS using the Quick-16S Library Prep kit and an Illumina (San Diego, CA, USA) sequencing platform.
Analysis of microbiota
Microbiota profiling was determined using the PrecisionBIOME™ bioinformatics analysis pipeline. Uclust was used to perform taxonomic classifications using a PrecisionBIOME™ custom proprietary database. Phylotypes were computed as percentage proportions based on the total number of sequences in each sample. Relative abundances of bacteria compared to fungi were determined, assuming an equivalency of one 16S rDNA copy to one fungal ITS copy. Absolute microbial quantification was achieved using a real-time PCR approach using primers targeting the V1–V3 and ITS regions for bacterial and fungal quantification, respectively. Species-level resolution of this sequencing approach was previously confirmed by shotgun sequencing (Alexis et al, 2021).
Statistical analysis
Unless otherwise stated, results were expressed as mean values with standard deviation. Measurements of α-diversity and evenness were calculated using the Shannon index, Simpson index, and the number of observed species. β-diversity was calculated using Bray-Curtis distance at the species taxonomic level for both bacteria and fungi, represented as principal coordinate analyses and statistically compared between groups using a Permutational Multivariate Analysis of Variance. Linear discriminant analysis and effect size (LEfSe) were used to identify taxa that were significantly enriched in each group using the default settings (QIIME version 1.9.1; P<0.05 was considered significant; Peer et al, 2022).
Analyses of variance and false discovery rate control to correct for type I errors were performed on the species-level relative abundance data of this analysis. Species with p<0.05 were considered significant. A presence-absence data matrix of species by site was generated by assuming species with abundance greater than 1% as present and less than 1% as absent. The “co-occur” function from R (“co-occur” package in R version 3.5.2, R Core Team, 2013) was used to generate pairwise classification of species having positive, negative, and random associations. The core microbiota was determined based on taxa detected with ≥5% relative abundance and in ≥50% of all samples. Following the sequencing of the samples collected and subsequent LEfSe, the relative abundance of different organisms present in the samples was analysed and compared between groups.
Results
Bacterial microbiota among those with diabetes and without diabetes
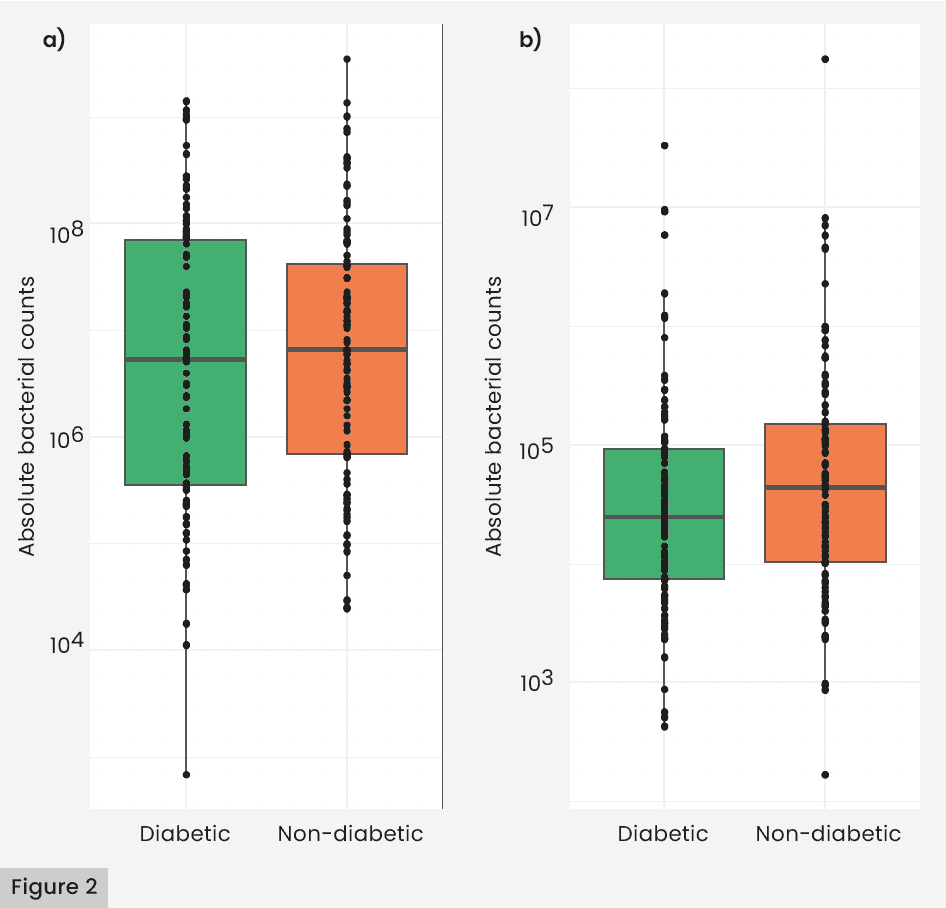
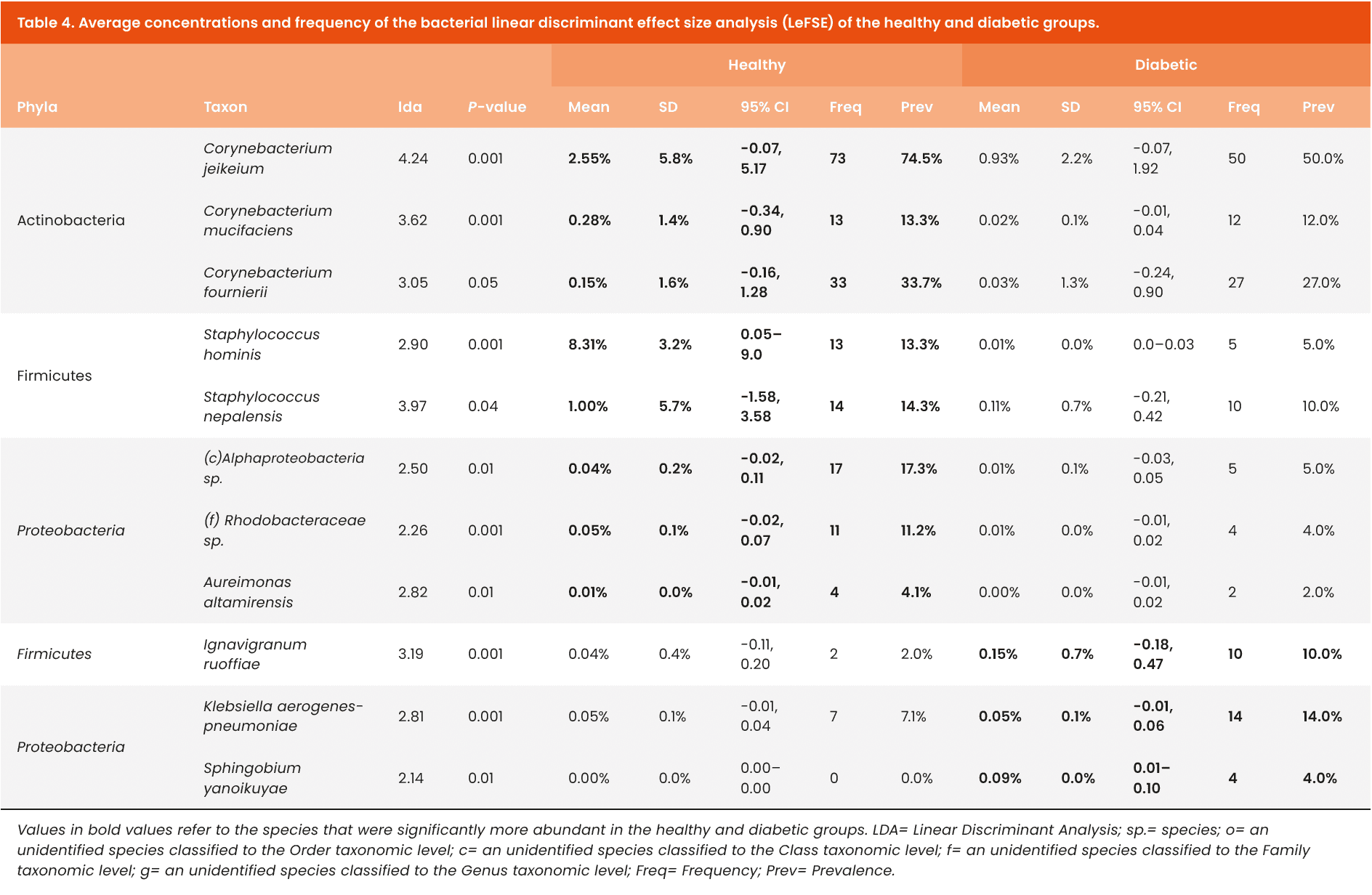
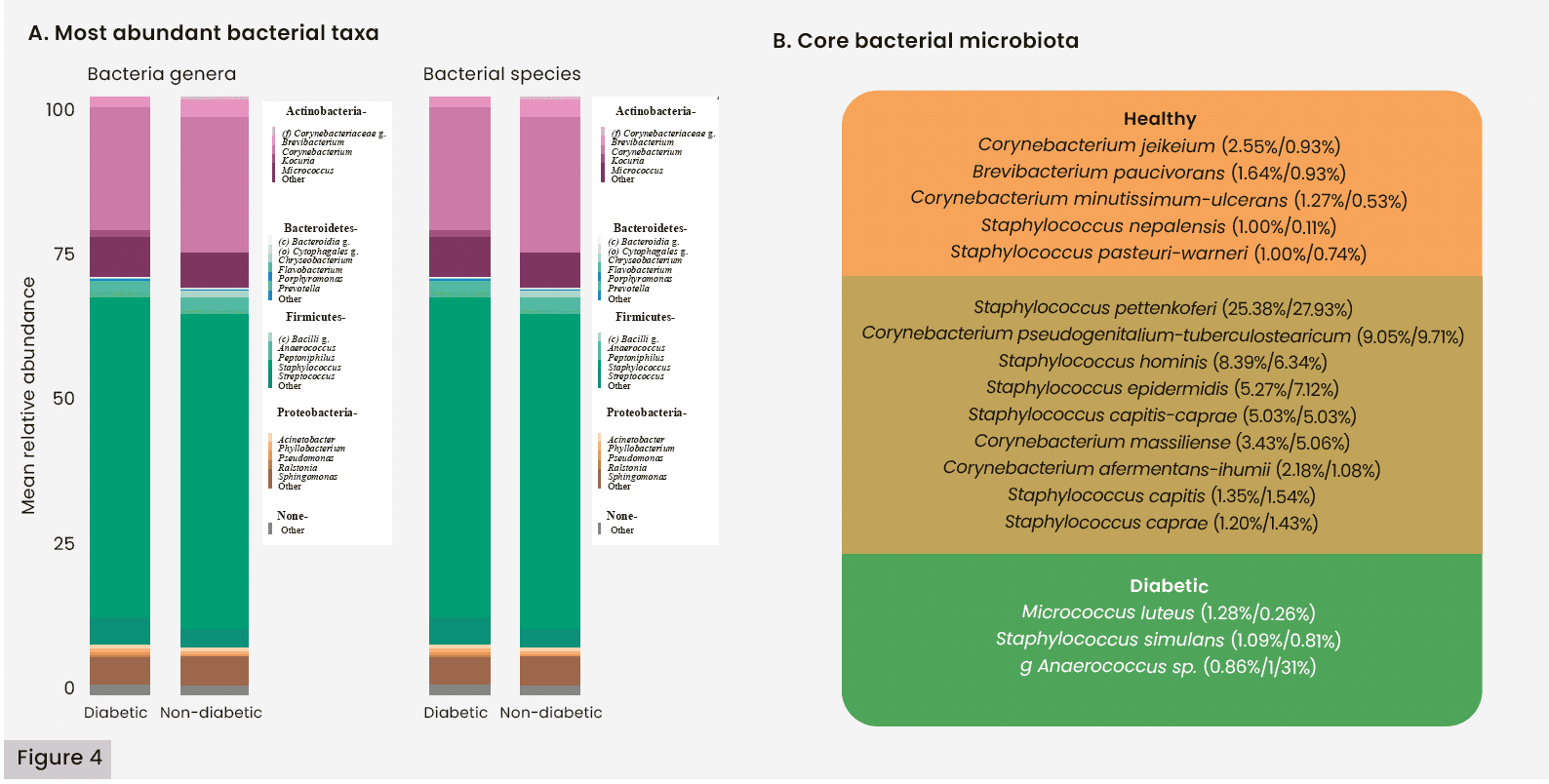
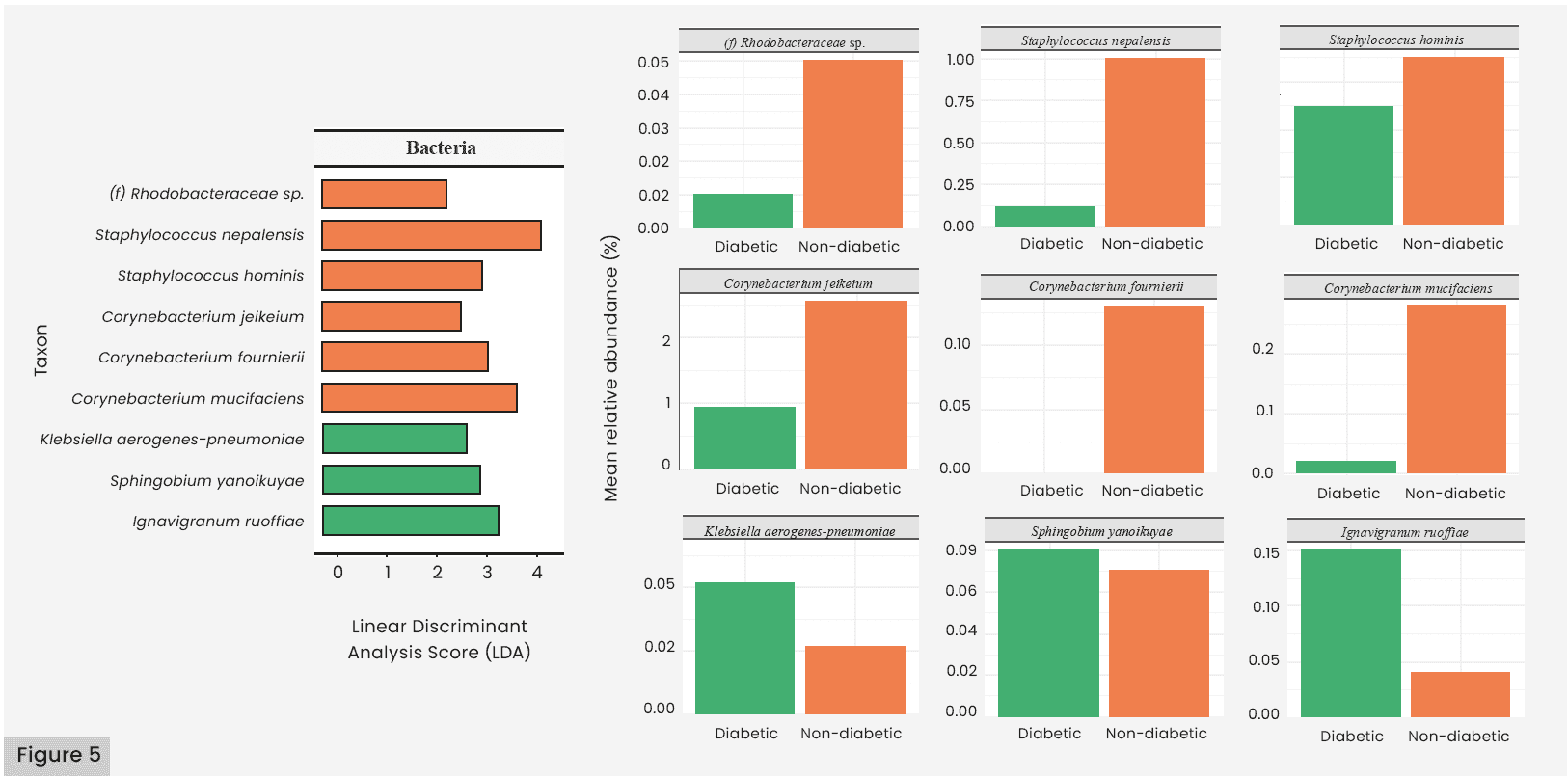
Overall, bacterial cell counts [Figure 2a]and bacterial diversity [Figure 3a–d] were similar between those with and without diabetes (p>0.05). As shown in Figure 4a, the most abundant bacterial species in both groups included taxa from the Actinobacteria phyla, such as species from the Corynebacterium genus (g-Corynebacterium spp.) and g-Micrococcus spp., and taxa from the Firmicutes phyla, such as multiple g-Staphylococcus spp. Figure 4B details the core bacterial microbiota. The Gram-positive bacteria Staphylococcus hominis and S. epidermidis were most abundant across both groups, while Corynebacterium jeikeum was distinct to the healthy group, and Micrococcus luteus was distinct to the diabetic group. Table 4 details the bacterial genera and species that were found to be significantly abundant in each group from the LEfSe Analysis. Figure 5 highlights taxa that were greater than 0.05% mean relative abundance. Among the significant species in the healthy group are three Corynebacterium genus (C. jeikeium, C. fournierii, and C. mucifaciens), two species from the Staphylococcus genus (S. hominis and S. nepalensis) and three from the Proteobacteria phyla, including an unidentified species from the Rhodobacteraceae family. In the diabetic group, Klebsiella aerogenes-pneumoniae, Sphingobium yanoikuyae, and Ignavigranum ruoffae were found to be significantly more abundant than in the non-diabetic group.
Fungal microbiota among those with diabetes and without diabetes
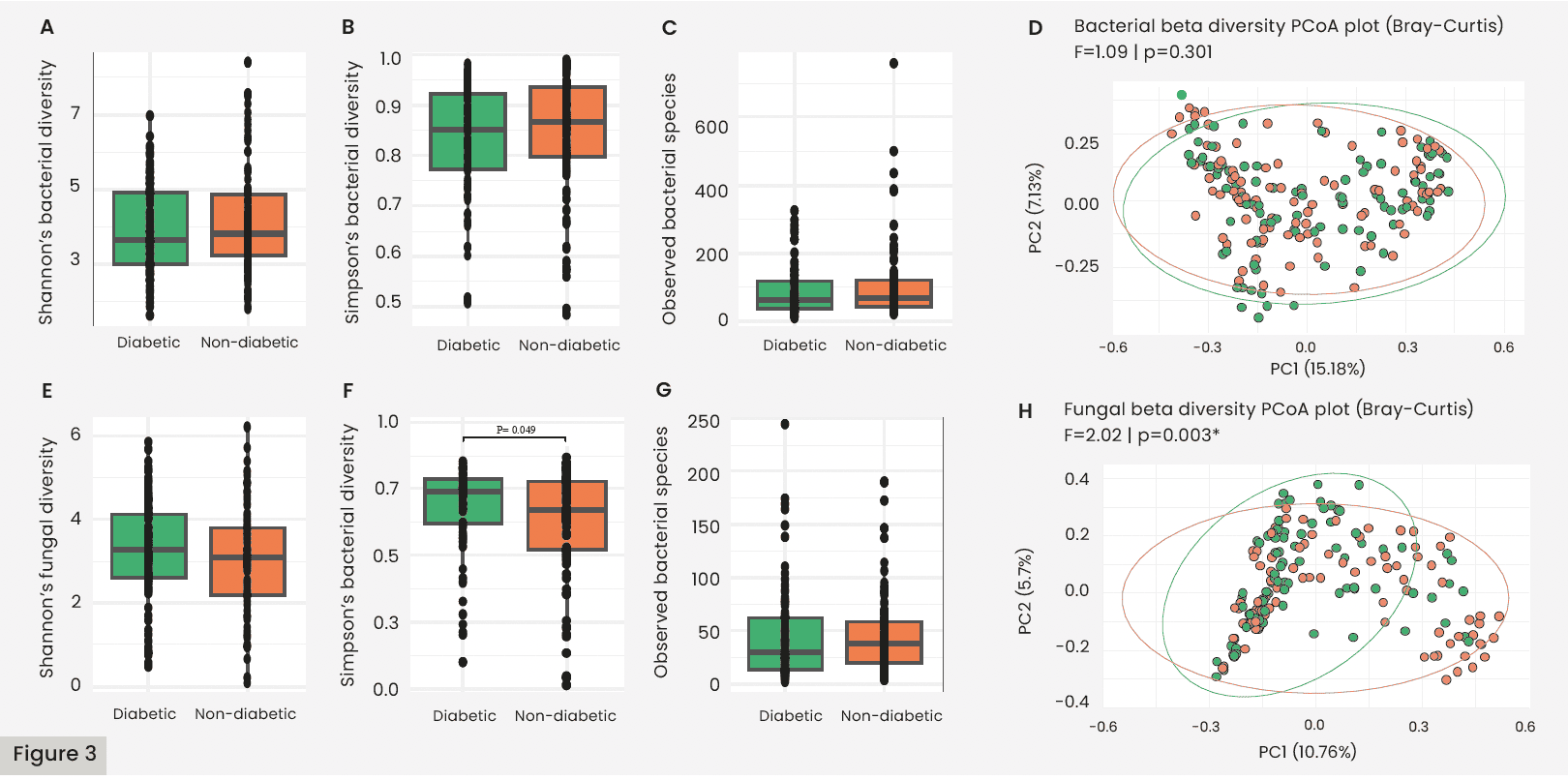
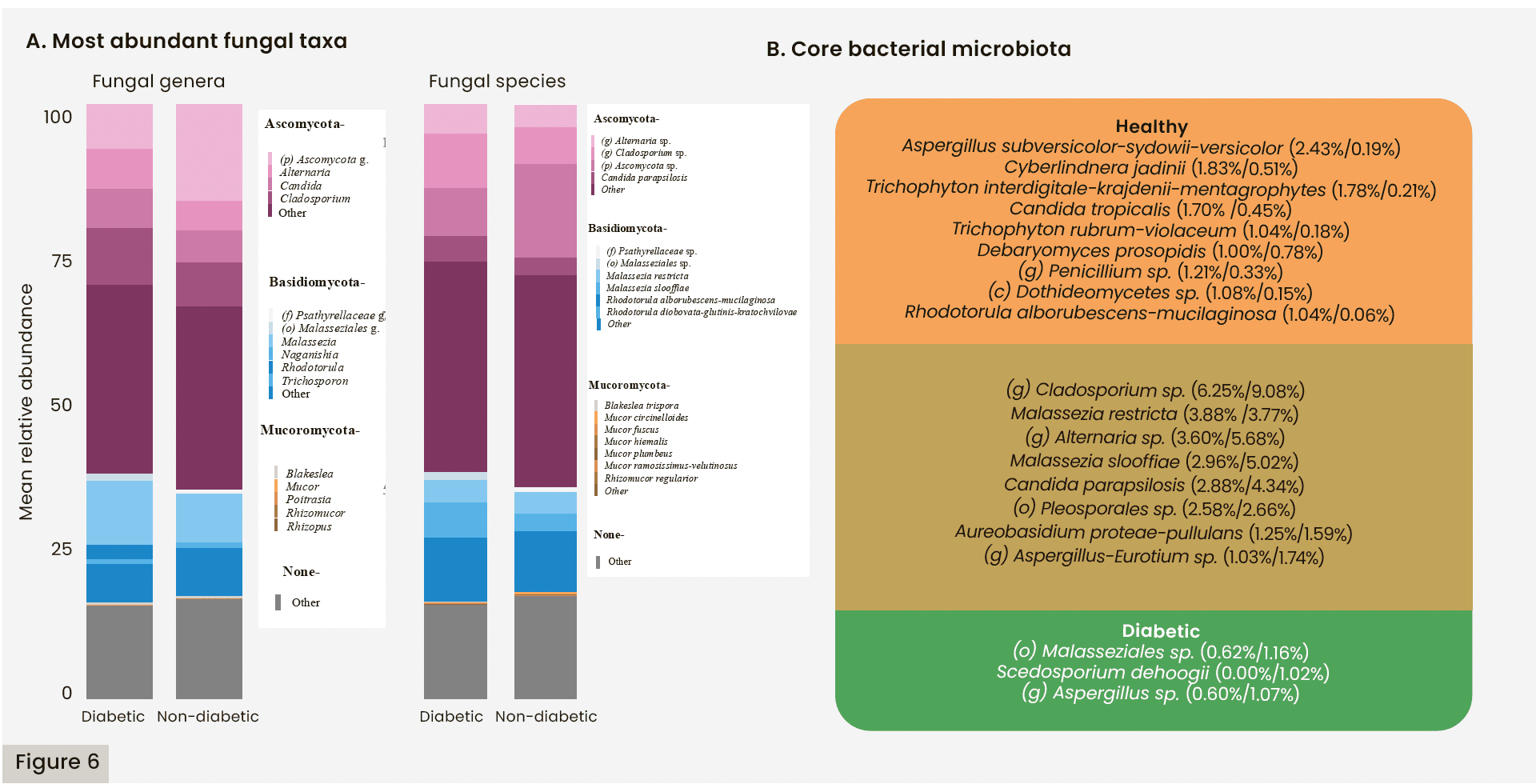
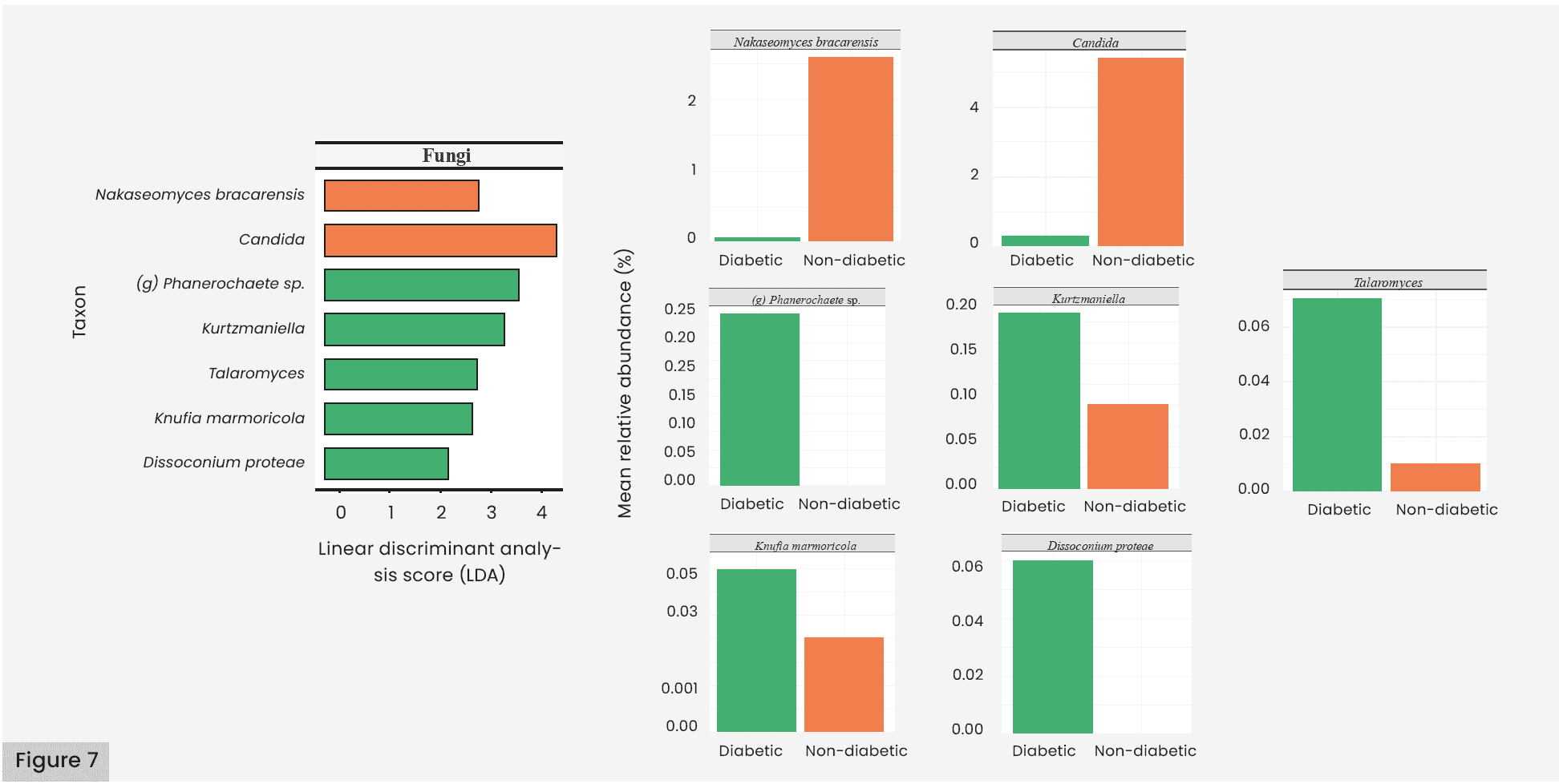
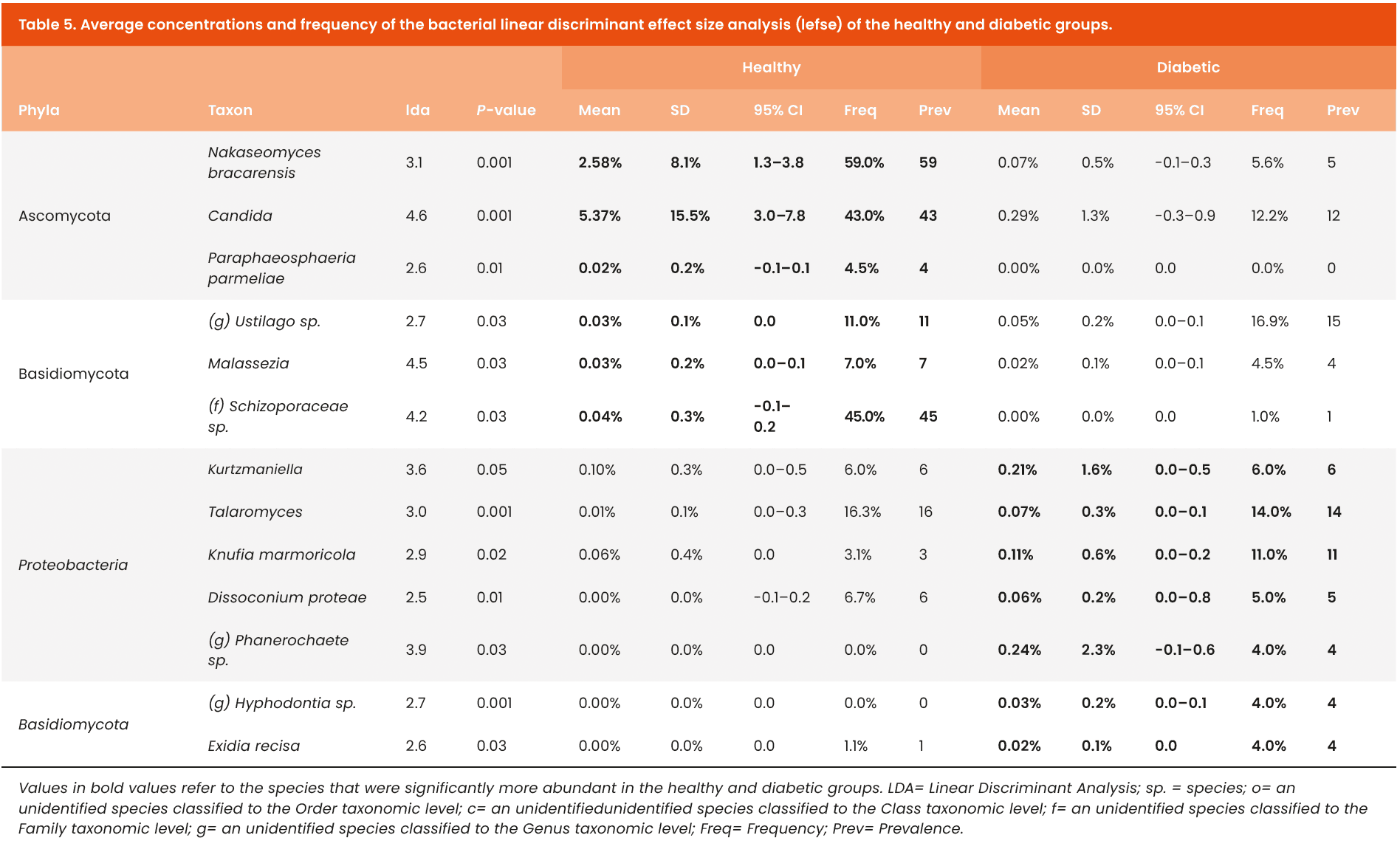
Fungal cell counts [Figure 2B] were similar between the diabetes and healthy groups. As shown in Figure 3E–H, Shannon’s Diversity and Observed Species were found to be similar between groups, but there was a significant difference between Simpson’s Fungal Diversity (p=0.049) and Bray-Curtis Beta Diversity (p=0.003). Figure 6A overviews the most abundant fungal species in both groups, which included taxa from the Ascomycota phylum, such as an unidentified species from the Alternaria genus (g-Alternaria spp.) and g-Candida spp., and taxa from the Basidiomycota phylum, such as g-Malassezia spp. Figure 6B details the core fungal microbiota. The shared microbiota included Malassezia restricta and g-Cladosporium sp. Among the fungal species distinctly identified in the healthy group were Candida tropicalis and Trichophyton rubrum. Table 5 details the fungal genera and species that were shown to be significantly abundant in each group from the LEfSe analysis. Figure 7 highlights fungal genera and species that were greater than 0.05% mean relative abundance. Among the healthy group, the Candida genus and Nakaseomyces bracarensis were significantly more abundant than in the diabetic group. In the diabetic group, (g) Phanerochaete sp., Kurtzmaniella, Talaromyces, Knufia marmoricola, and Dissoconium proteae were more abundant than in the non-diabetic group.
Co-occurrence interactions between microbes among diabetic versus healthy Individuals
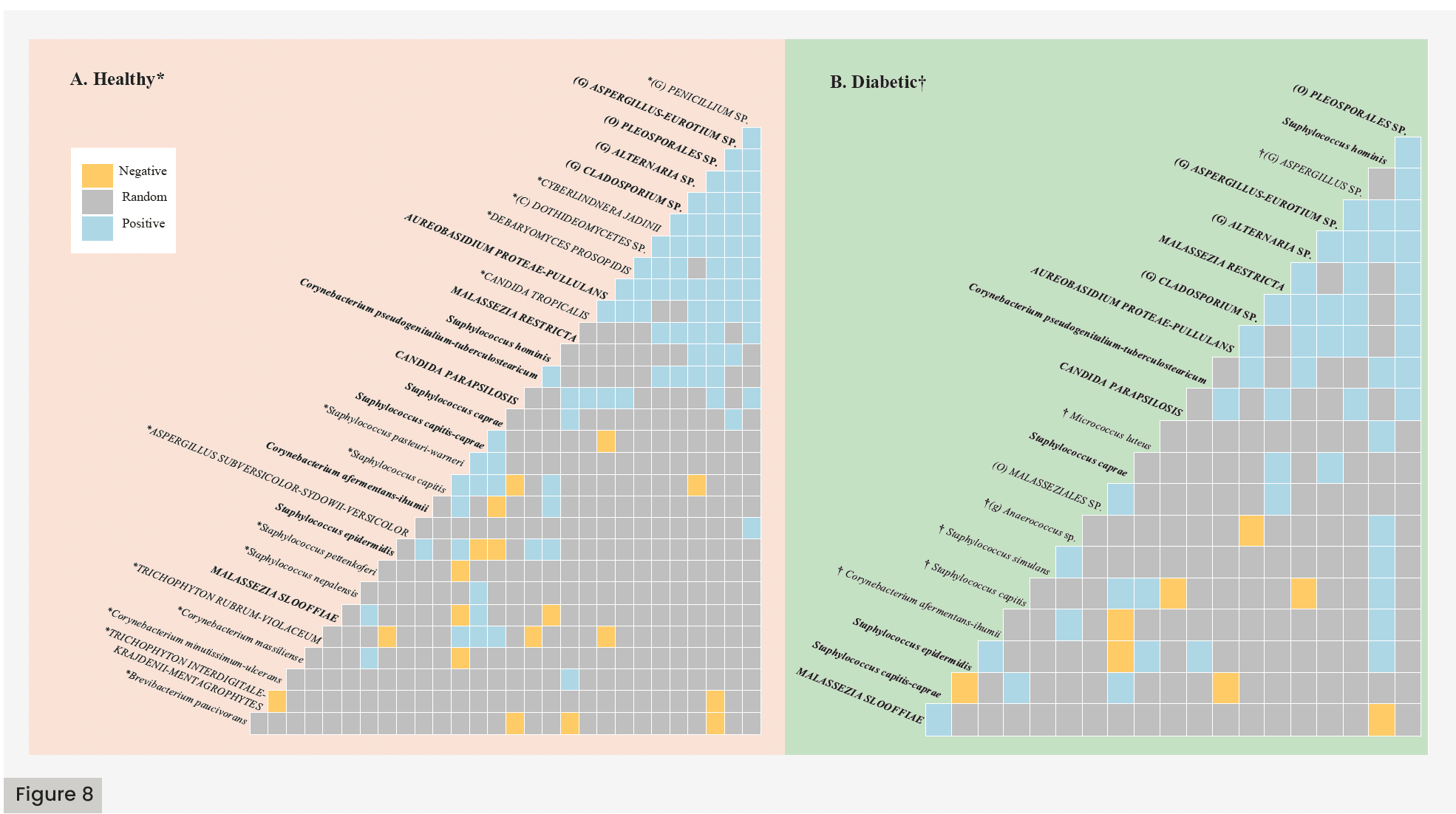
A co-occurrence analysis was conducted among the top microbial species in the healthy group [Figure 8A] and diabetic group [Figure 8B] to identify interactions between bacterial and fungal species within a shared environment to determine whether the presence of a pathogen may be influencing commensals in response to diabetes. In both groups, there were several positive interactions between similar fungal species, including unidentified species from the Aspergillus-Eurotium, Alternaria, and Cladosporium genera, an unidentified species within the Pleosporales order, as well as Aureobasidium proteae-pullulans and Malassezia restricta. However, in the healthy group, this cluster of interactions included more microbes: an unidentified species from the Penicillium genus, Cyberlindnera jadnii, an unidentified species from the Dothideomycetes class, Debaryomyces prosopidis, and Candida tropicalis. In the diabetic group, this cluster included an unidentified species from the Aspergillus genus. Although Staphylococcus (S.) hominis was present and interacted with several microbes in both groups, there were shown to be more positive interactions between other microbes in the diabetic group versus the healthy group. Nine positive interactions were seen in the diabetic group (e.g. (g) Aspergillius sp., (g) Alternaria sp., S. simulans, and S. captis) and only four positive interactions were seen in the healthy group (e.g., S. captis and S. epidermis). Both the healthy and diabetic groups showed a negative interaction between S. hominis and the fungal species, Malssezzia sloofiae. Lastly, both groups showed a positive interaction between Corynebacterium pseudogenitalium-tuberculostearicum and S. epidermis.
The differences in bacteria and fungi between healthy and diabetic skin sample locations
Supplementary Figures 1 and 2 highlight some of the most significant bacteria and fungi when comparing skin sites and health status from an additional LeFSe Analysis. In the Plantar-Healthy group, two species from the Staphylococcus genus (S. hominis and S. arlettae), among others (Supplementary Figure 1). The Gram-positive bacilli Cutibacterium acnes (previously named Propionibacterium acnes) was among the most significant bacteria in the Plantar-Diabetes group. In the IDS skin sites, the healthy group had more abundant Staphylococcus devriesei and two species from the Corynebacterium genus (C. jeikeium and an unidentified Corynebacterium species), whereas Brevibacterium linens was significantly more abundant in the diabetic group. Among fungal species, Perenniporia subacida and Alternaria alternata-cerasi-ellisii species were significantly abundant in the Plantar-Healthy group, and a (g) Paraphoma sp. was more significantly abundant in the Plantar-Diabetes group. Among the IDS site groups, Malassezia restricta was more abundant in the healthy group, and the Malassezia genus was more abundant in the diabetic group.
The differences in bacteria and fungi between males and females
Supplementary Figures 3 and 4 highlight the most significant bacteria and fungi when comparing biological sex. Among the healthy group, two species from the Corynebacterium genus (C. jeikeium and C. singular) and two from the Staphylococcus genus (S. felis and S. arlettae) were more abundant in males, and the Actinoplanes genus and an unidentified species from the Solirubrobacter genus were among the more abundant in females. In the diabetic group, Ignavigramum ruoffiae and an unidentified species from the Anaerococcus genus were more abundant in males, while the Lactobacillus genus was more abundant in females. Among fungal taxa, Knufia marmoricola and (g) Hyphodontia sp. were some of the more abundant fungi in the healthy male group, where no taxa were greater than 0.05% mean relative abundance and significant in the healthy females. Among the diabetic group, Malassezia and Candida genera were more abundant in males, and an unidentified species from the Schizoporaceae species was more abundant.
The differences in bacteria and fungi between participants across the Fitzpatrick skin type scale
Supplementary Figures 5 and 6 highlight the most significant bacteria and fungi among participants along the Fitzpatrick skin type scale. Bacteria that were most abundant in the fairest skin tone, level I, include Staphylococcus pasteuri-warneri and Corynebacterium jeikeium. In the level II group, Anaerococcus procencensis and the Peptoniphilus genus were among those most significantly abundant. Participants with the type III skin tone showed significant abundances of bacteria like Corynebacterium resistens and Corynebacterium imitans. In the type IV group, Staphylococcus pettenkoferi was significantly abundant, whereas an unidentified genus in the Saccharibacteria phyla was more abundant in the type V group. In the type VI group, Staphylococcus petrasii and Corynebacterium afermentans-ihumii were more abundant. Several fungal taxa that were found to be significant in each group included two Malassezia species in the type I group, Aspergillus penicillioides in the type II group, an unidentified Cladosporium genus in the type III group, the Candida genus in the type IV group, Aspergillus gracilis in the type V group, and an unidentified species in the Peniophora genus in the type VI group, among others.
Discussion
This study on skin microbiota of the foot revealed compelling patterns among healthy individuals and those with diabetes that may provide insight into possible therapeutic modalities in the future. When considering the microbiota as a whole, the similarity in bacterial cell count and diversity seen in this study provides important insights that the skin microbiota may be resilient enough to maintain its overall integrity in those with diabetes.
As it pertains to the fungal microbiota, the differences in diversity suggest that diabetes may impact the fungal integrity of the microbiota. In addition, the individual bacterial and fungal differences seen between healthy individuals and those with diabetes point to the plausibility that particular microbes may distinguish diabetes. For instance, Staphylococcus and Corynebacterium species were more abundant in the healthy group, whereas Klebsiella and Sphingobium species were more abundant in the diabetes group.
Typically, the commensal skin bacterial microbiota is comprised of Staphylococcus, Cutibacterium, and Corynebacterium species. In confirmation, the present study identified two Staphylococcus species (S. homini and S. nepalensis) and three Corynebacterium species (C. jeikeium, C. fournierii, and C. mucifaciens) to be significantly more abundant in the healthy group. In a diabetic population, Gram-negative bacteria, including (g) Klebsiella and (g) Corynebacterium species, comprise the skin microbiota and have been linked to diabetic complications (i.e. diabetic foot; Heravi et al, 2019; Zhang et al, 2023).
In the current study, one Gram-negative bacterial species, Klebsiella aerogenes-pneumoniae, was significantly more abundant in the diabetic group. Still, some bacterial species were found to be more abundant in the diabetic group that have not yet been identified in a dysbiotic skin microbiota, Sphingobium yanoikuyae and Ignavigranum ruoffiae. Given that this is one of the seminal studies looking at diabetic skin microbiota, more evidence is needed to elucidate the role of these microbes in diabetes.
Among fungal taxa, previous studies have linked an increase in candidiasis in people with diabetes, with speculated reasons being the immune-compromised status or increased blood sugar levels in these groups (Mohammed et al, 2021). In confirmation with this evidence, the Candida genus was found to be more abundant in the healthy group, particularly healthy females, compared to diabetic individuals. However, it is important to note that the Candida genus comprises both commensal and opportunistic species. Since the current analysis did not identify any particular species that has previously been shown to be opportunistic in diabetic patients, such as Candida albicans (Shahabudin et al, 2024), the implications of these findings are unclear and further investigation is necessary.
While some of the fungal species identified in the diabetic group have not previously been identified in the skin microbiota of a diabetic population, several fungal species shown to be significantly abundant in the IDS-Diabetic and Plantar-Diabetic groups are consistent with previous evidence. For instance, Malassezia was higher in the IDS-Diabetic group, a fungus that has previously been attributed to dysbiosis in the skin (Saunte et al, 2020). In addition, both Malassezia and Candida were shown to be more abundant in the diabetic group when stratifying for biological sex. As such, there may be a biological sex-dependent response to the skin among those with diabetes. Interestingly, two Candida species (C. tropicalis and C. parapsilosis) were present in the healthy group, but only C. parapsilosis was present in the diabetic group. In addition, similar Malassezia species (M. restrica and M. sloofiae) were seen in both groups.
Previous research has highlighted the importance of bacterial-fungal relationships in health and disease. For example, the virulence of Staphylococcus aureus has been shown to be reduced when exposed to the commensal Corynebacterium species Ramsey et al (2016). Fungi isolates tend to take advantage of reduced bacterial levels following antibiotic therapy (Kalan et al, 2016; Drummond et al, 2022) and also levels can be higher in immunocompromised individuals, such as those with diabetes, as identified by Shahabudin et al (2024). These results suggest a possible risk factor for diabetes patients, particularly as fungal infections can be difficult to manage effectively due to the increased kill time required, combined with the growing challenge of antifungal resistance (McCormick and Ghannoum, 2024). In the current study, among the bacterial and fungal interactions observed in the co-occurrence analysis, both groups showed a similar cluster of positive interactions between mostly fungi, while the positive interaction cluster in the healthy group included more commensal fungi than in the diabetes group, including Penicillium and Staphylococcus species. These findings, in conjunction with the lower fungal diversity seen in the diabetic group, suggest that, in a diabetic environment, there may be fewer commensal fungi present to maintain the diversity and microbial integrity of the microbiota in those with diabetes. Notably, the diabetic group showed more positive interactions between the commensal, Staphylococcus hominis, and several microbes. These interactions may be occurring in an attempt to counteract disease-related perturbations to the microbiota and maintain the microbial integrity of a skin microbiota that is compromised, like that in a diabetic environment. For instance, co-culture experiments of Cutibacterium acnes with S. epidermidis have reported a significantly enhanced biofilm formation for C. acnes in both aerobic and anaerobic conditions (Kaplan, 2023), which may be implicated in greater skin pathogenesis. In contrast, there were fewer interactions with S. hominis in the healthy group, suggesting less of a need for S. hominis to counteract any pathogens that may be present. Still, these findings must be interpreted with caution, as there is limited evidence on the mechanism or involvement of such microbial interactions in diabetes.
There were also several differences in Staphylococcus species seen across skin tone types (Fitzpatrick scale I–VI), which warrant future consideration to analyse both different ethnicities to determine differences in diabetic responses, as well as more severe cases of diabetes in which the skin may change colour or integrity due to diabetic wounds. It is possible that the differences seen in the microbiota could be linked to moisture levels of the skin, with dark skin shown to have lower basal water content in the stratum corneum because of differences in trans-epidermal moisture loss (Wan et al, 2014; Alexis et al, 2021; Peer et al, 2022), which in turn could affect the microbiota composition. However, to our knowledge, this was the first study to consider skin tone differences in a diabetic population. Given the paucity of research into the skin microflora in different skin tones, more evidence is needed that explores these skin microbial differences further.
While the findings from this study contribute to research in the diabetic skin microbiota, some limitations must be addressed. First, this study was conducted at a single site and at a single time point; therefore, future studies should include specimen collection and results from multiple centres to encompass any regional differences in the skin microbiota that may be seen. Second, longitudinal measures of skin flora and antimicrobial medication may help track any shifts in populations, especially if ulceration occurs, to further understand any links between skin flora and diabetic foot infection. Higher levels of Gram-negative bacterial species may be a result of antibiotic treatments for previous infections, which can wipe out the healthy cutaneous microbiota (Jo et al, 2021). This pattern is concerning as infections caused by Gram-negative organisms are increasingly becoming some of the most difficult to treat due to increasing antimicrobial resistance in this group (Russo et al, 2022). These organisms are also prolific biofilm formers, further adding to their tolerance to antimicrobial intervention (Pompilio et al, 2021; Zhao et al, 2023). Antimicrobial resistance was not assessed in this study; however, future investigation is warranted.
Conclusion
The findings of this study underscore the intricate nature of the skin microbiota, influenced by factors such as geographic location, comorbidities, such as diabetes, biological sex and skin type. By taking these variables into account in future research, it will be possible to better identify the risks involving the microbiota that are associated with diabetic foot infections following ulceration. This insight will be invaluable in crafting targeted hygiene practices that promote and preserve healthy skin microbiota.
Supplementary figures can be accessed here.
